Home / Work Packages
Work Packages
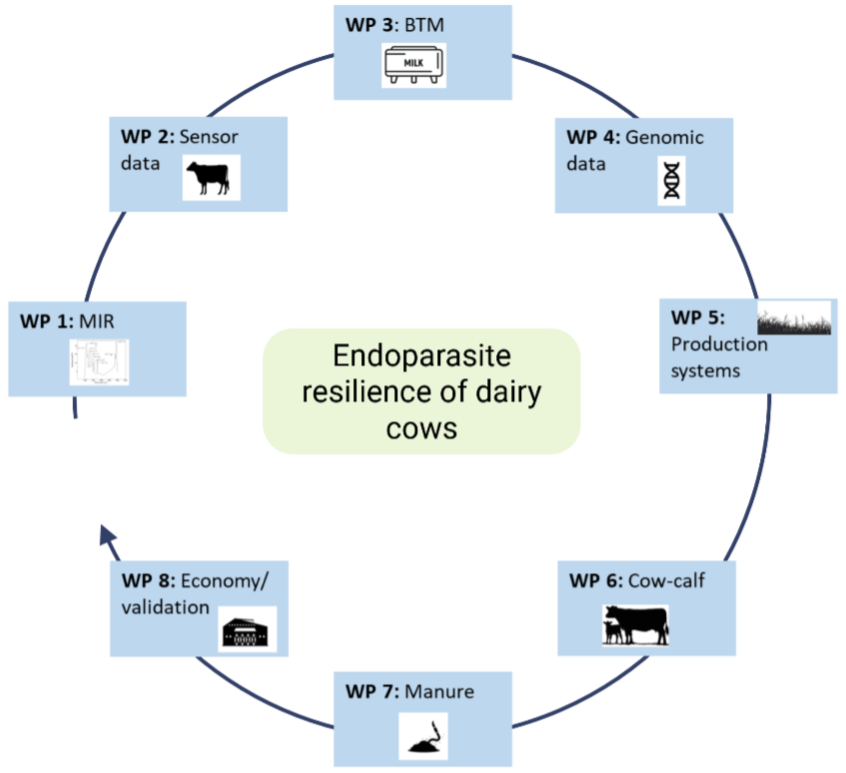
Work Packages overview (quick navigation)
| WP | Title |
|---|---|
| WP1 | Milk MIR-based detection of endoparasite burden |
| WP2 | Sensor-based monitoring and early warning systems |
| WP3 | Bulk tank milk antibody diagnostics |
| WP4 | Genomic tools for parasite resistance |
| WP5 | Climate, pasture and production system effects |
| WP6 | Milk quality and calf health |
| WP7 | Smart manure management |
| WP8 | Validation and economic evaluation |
| WP9 | Project management and communication |
General work package description
TechGen4Health is divided into 9 work packages WP1-9. All partners will follow their expertise, supported with data and methods from the partner countries.
In WP1, ULiège will develop and apply dedicated milk MIR spectra-based prediction equations to detect burden of endoparasites, which can then be used afterwards by farmers and public services to obtain information about the prevalence and infection status of individual cows and at herd level.
In WP2, KUL will monitor endoparasite infections in grazing cattle using on-farm sensor technology and quantify the disruption to performance, health and welfare caused by these infections. Digital tools for early warning and to analyse disease development and recovery will be developed.
In WP3, UCPH explores and validates BTM-Ab diagnostics as a tool for endoparasite control, correlates the occurrence with drug use, and analyses seasonal dynamics and changes associated with preventive strategies.
In WP4, JLU improves the endoparasite resistance in grazing herds using genomic selection tools. In this regard, the developed genomic herd management tool enables the early selection of parasite resistant female calves being later on parasite resistant grazing cows with a high welfare and health status.
In WP5, UBO investigates the impact of climate, grassland parameters and production systems on endoparasite burden of dairy cows. Grassland parameters will be characterized using drones and satellites, and climate will be measured continuously with sensors, aiming on early prediction of endoparasitic infections based on dense environmental descriptors.
In WP6, UL investigates the effect of endoparasite infections of cows on milk quality and on the calf health. Differences between breeds, dairy/suckler cows, calf-rearing types and types of pasture will be analysed for infection status and milk quality of cows, and associated with calf health.
In WP7, WLR analyses the effect of smart manure management on endoparasite spread and will develop an automated faecal egg counting method (AFEC) as a future substitute for the time-consuming lab methods. WLR will elaborate manure handling guidelines, to minimize parasite transmission risks.
In WP8, LSMU develops the methodological framework for comprehensive tool validations and overall economic evaluations. A cost-benefit analysis will be applied to develop guidelines for the integration of validated tools into regular farm practices. The final evaluation of applicability will be done in research herds from all partner countries. Hence, practical cattle farmers are directly involved in the disease prevention tool applications, strongly contributing to disseminations in grazing production systems.
Additionally, WP9 focusses on the project organisation, communication and the establishment of a data management platform. Regular video conferences and project meetings will be established to discuss project progresses, interim results and problems in detail. In addition, LSMU and JLU have created this centralised TechGen4Health homepage. A data management platform is also being established, where data from all partners will be stored centrally and made available to all participating scientists and associated partners.
Individual WP descriptions
WP1
WP1 aims to develop and apply dedicated milk mid-infrared (MIR) spectra-based prediction equations to detect burden of endoparasites as the core of alert and surveillance systems for the detection of endoparasite infections. We will provide a tool to inform farmers, but also public services on the in-farm prevalence and infection status of individual cows (MIR from milk recording) as well as at the herd level (MIR from bulk milk).
P2 will compile reference data sets for MIR analysis and establish the use of MIR for monitoring endoparasite infections for all partners. The transferability of the equations will be ensured through the standardisation of MIR. The equations will be validated and used throughout the partnership. The reference data sets will be expanded to include sensor data (WP2), milk antibody analyses (WP3) and other technologies.
WP2
WP2 aims to monitor endoparasite infections in grazing cattle using on-farm sensor technology. To this end, we will characterise normal (in-control) patterns and variation in the collected sensor time series to quantify perturbations (out-of-control) in performance, health, and welfare due to these infections. Ultimately, we target the development of digital tools for early warning alerts, and to analyse disease development and recovery, integrated with the outcomes of the other project WPs.
On-farm sensor time series of grazing cattle will be collected and processed to identify perturbations linked to endoparasite infections. This will result in digital tools to detect infections, predict their severity, and monitor recovery. These tools will aid in phenotyping resilient animals (link with WP4), assessing infection pressure (link with WPs 1, 3, 5, 7), and evaluating the economic impact (link with WP8) of cattle endoparasites.
WP3
Bulk tank milk-antibody (BTM-Ab) tests have been developed for liver fluke, lungworm and gastrointestinal nematodes (GIN) infection predictions based on Enzyme-linked Immunosorbent (ELISA) analyses. However, it is not fully clarified how to best interpret and act on the obtained information. WP3 aims to explore and validate BTM-Ab diagnostics as a tool for endoparasite control, and correlates the occurrence with drug use, as well as analyses seasonal dynamics and changes associated with preventive strategies in cow herds.
UCPH investigates BTM-Ab tests to evaluate presence of endoparasite infections in dairy herds focusing on liver and rumen flukes, lungworms, and GIN. UCPH carries out BTM-Ab tests and validates the results at herd level through individual recordings. Disease recordings and drug use will be analysed in relation to BTM-Ab at herd level, as well as BTM-Ab levels in relation to season and preventive strategies. BTM-Ab tests will be transferred to routine tests, even in partner countries.
WP4
WP4 aims on the improvement of endoparasite resistance in grazing herds using genomic selection tools. UGI will develop genomic herd management instruments, enabling the early selection of parasite resistant female calves to produce parasite resistant grazing cows with a high welfare and health status.
UGI will enlarge existing datasets by continuing faecal sampling for endoparasite traits in Germany, supported by respective activities in all partner countries. Based on extreme phenotypes for endoparasites, we select the most susceptible and most resistant cows for whole genome sequencing. The SNP genotypes of the remaining cows with endoparasite phenotypes will be imputed to sequence level. For a subset of cows with most extreme phenotypes for infections, endoparasites will be genotyped, to infer host-parasite interactions. Genomic predictions will be enhanced considering milk spectra (WP1), milk antibodies (WP3), sensor traits (WP2), milk composition (WP6), and inferred grazing production system characteristics (WP5).
WP5
WP5 aims to investigate the impact of climate, grassland parameters and production systems on endoparasite burden of dairy cows.
UBO will collect health and endoparasite parameters in a research farm in South Tyrol with two different production systems. Using satellites and drones, grassland parameters like pasture condition and grazing intensity, are determined. Climate data will be continuously obtained using sensors and weather station data. The aim is to find solutions for the control of endoparasite infections through system improvements based on dense ‘environmental data’. Including e.g. milk-based parameters according to the partner’s WP, production system specific early prediction models for the control of endoparasites will be developed. Herd infection risks will be modelled based on the environmental data.
WP6
WP6 aims to investigate the effect of cow’s health status on milk quality and, from a transgenerational perspective, on the calf health status.
UL uses different breeds, types of pasture (intensive, extensive, mountainous), and calf-rearing forms including mother-bonded rearing. This enables analyses of direct correlations between cow’s infection pressure on milk quality and on calf health. Cow-calf pairs will be used for monitoring of health traits across generations. Analyses of the lactoferrin-content via MIR (WP1) and of milk antibodies (WP3) lead to connections with partners. Digital tools of KUL will be used to monitor calf health.
In consequence, structural equation models will be applied to study the time-lagged pathway of cow’s infection at cow milk quality and at calf health and weight. Models will be applied in the different grazing systems in Slovenia, and extended to the other partner countries
WP7
WP7 aims to reduce endoparasite infections through smart manure management. As manure serves as carrier for parasites and their eggs, WP7 will create a theoretical framework that models the spread of parasites through manure handling. The infection pressure of parasites will be analysed for manure with and without the use of modern handling techniques like emission-reducing floors, faeces-urine separation, acidification, additives and composting.
Automated faecal egg counting methods (AFEC), using artificial intelligence (AI) and automatic vision techniques, will be developed and validated with faecal lab analyses of UGI. AFEC will be used for a comprehensive assessment of the endoparasitic load of manure on floors, in storage, digesters, as RENURE (REcovered Nitrogen from manURE), and applied to land.
Combined with the results of the partners, production system specific predictions and farm specific prevention strategies will be developed and shared with the stakeholders. WLR generates manure handling guidelines to minimize parasite transmission risks.
WP8
The aim of WP8 is the validation and economic evaluation of the preventive health management tools and the practical implementation of dissemination activities. A pilot system for validation and economic evaluation of preventive health management tools from all partners will be implemented on LSMU Baisogala AHC dairy research farm. The results will be decisive for the future adoption and integration of these tools into routine dairy farming processes. An economic evaluation of the health management tools for detecting infections in cattle herds will be conducted. The evaluation compares the costs and benefits of these tools, assessing their impact on herd health, productivity, and overall farm profitability by applying a cost-benefit analysis model.
Subsequently, the predictions and technical implementations developed by LSMU will be finally evaluated for applicability in selected research herds in the partner countries. Dissemination will promote the adoption of the proposed innovative tools by sharing them with stakeholders, including farmers, professionals, and policymakers. Workshops will be organised in each country.
WP9
WP9 focusses on the project organisation (e.g. regular video and face-to-face meetings), the establishment of a data management platform and this homepage here. Regular video conferences and project meetings will be used to discuss project progress, interim results and problems in detail.
